Informatics of docking libraries & docking screens
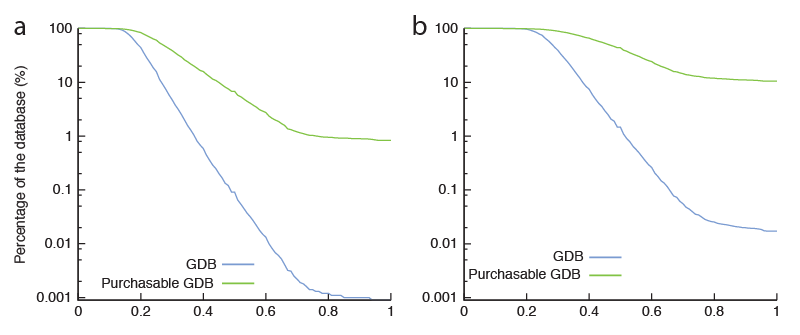
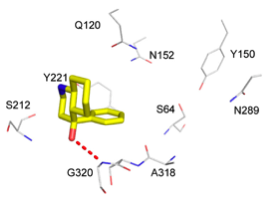
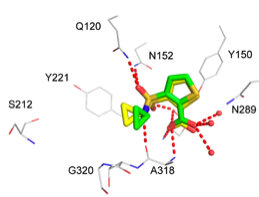
Comparing the potential size of chemical space to the actual size of screening libraries (Hann, 2001), it is a miracle that screening ever works. One reason why it does, we have argued (Hert, 2009; Figure 1), is that the libraries that are actually screened are fortuitously biased towards bio-relevant chemotypes. The chemical content of our libraries, either for high-throughput, fragment, DNA-encoded, or docking screens, is as important for the success of a discovery campaign as is the quality of our assays and the quality of our scoring functions.
An active project in the lab is to understand how to enrich libraries with bio-relevant molecules, and with such molecules grow the libraries. We have investigated how docking compares to high-throughput screens (HTS) against targets such as PTP-1B (Doman et al, 2002), AmpC (Feng et al, 2007), β-lactamase (Babaoglu et al, 2008), and Cruzain (Ferreira et al, 2010), and how docking fragments compares to docking larger, lead-like molecules, again using AmpC (Teotico et al, 2009) and TEM-1 (Chen et al, 2009) as model systems. A very recent direction involves efforts to grow our docking libraries by log orders. Already, at 3 to 6 million molecules, docking screens are larger than most HTS campaigns; over the next several years we expect the docking libraries to expand by 1000-fold, introducing fascinating new challenges and huge opportunities.
A close coupling between theory, docking, and experimental testing is central to the enterprise. Key papers include:
- S Barelier et al. Increasing chemical space coverage by combining empirical and computational fragment screens. ACS Chem Biol 9, 1528-35 (2014).
- RS Ferreira et al. Complementarity between a docking and a high-throughput screen in discovering new cruzain inhibitors. J Med Chem. 53, 4891-905 (2010).
- DG Teotico et al. Docking for fragment inhibitors of AmpC beta-lactamase. PNAS 106, 7455-60 (2009).
- J Hert et al., Quantifying biogenic bias in screening libraries. Nature Chem Biol 5, 479-83 (2009).
Supported by NIGMS GM59957 & GM71896.